Covalent stapling strategies that stabilize a particular secondary structure have garnered much attention as interest in peptide therapeutics continues to grow. One such strategy - using olefin-bearing unnatural amino acids covalently bonded using ring-closing metathesis - has been exploited to the greatest extent thusfar.
In today's post, I'll discuss some strategies to overcome DMF poisoning of the Grubbs catalyst used during the metathesis reaction towards fully automating the synthesis and secondary chemistry required for stapled peptides.
As many of you know by now, I think a lot about automating any and all chemistry steps that I can during my peptide synthesis. While this strategy serves primarily to free up my hands-on energy and time, it often requires some additional optimization before fully implementing a fully automated chemical process. In a previous post, I evaluated some conditions that can be used to expedite the ring closing metathesis reaction for stapled peptide synthesis, but these were undertaken with manual addition of the reagents.
It seems a little crazy to think that adding a reagent with a robot liquid handler could impact the outcome of a reaction, but it can! So whenever I undertake some new chemistry, particularly with metal catalysts, I manually add the reagents.
Unfortunately, I learned the above mentioned fact the hard way. No matter which of the conditions I tried while evaluating different microwave RCM reactions the best conversion efficiency I observed was about 50 percent when allowing the liquid handler to add my Grubbs catalyst solution, Figure 1. 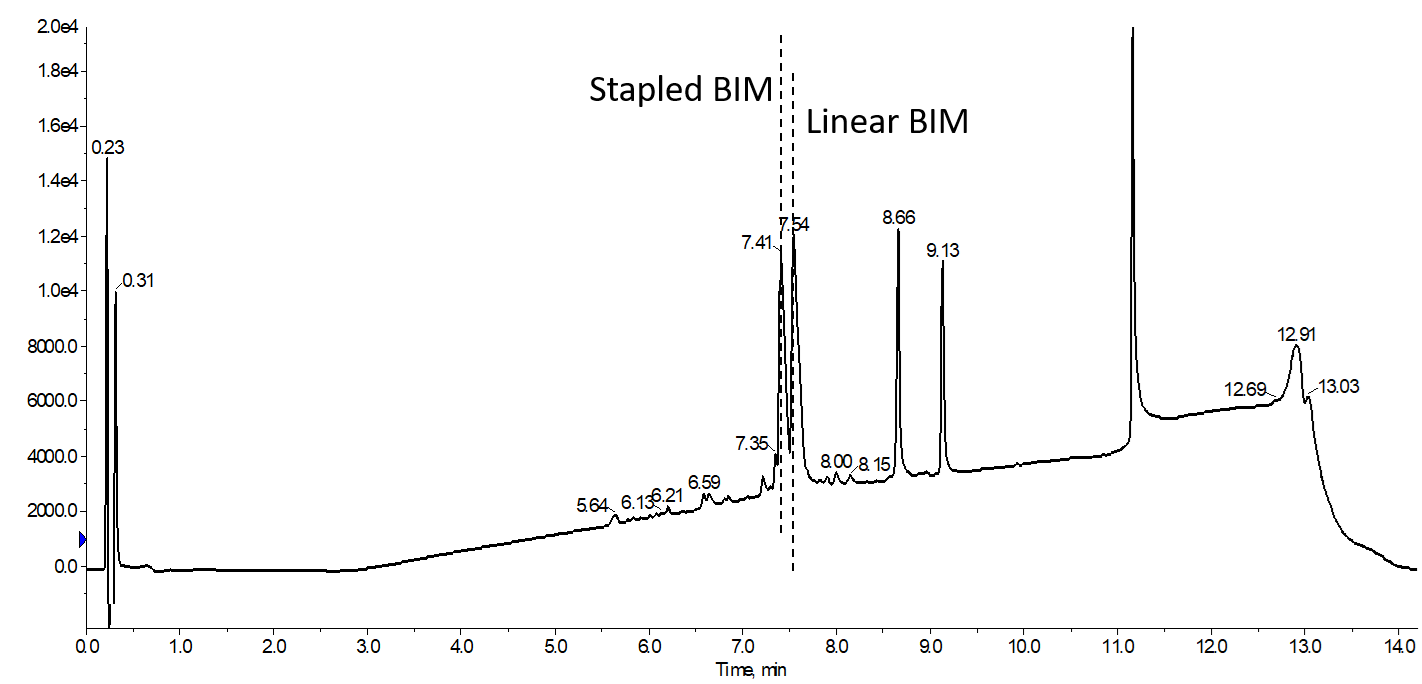
Figure 1. Conversion of linear BIM to the desired stapled BIM peptide product. The metathesis reagents were added automatically added with the robot liquid handler and reaction allowed to proceed for 60 min at 40°C, conditions that enabled complete conversion to the desired product with the reagents added manually.
Now, for a reaction that I know can be driven to completion, a 50% conversion is simply unacceptable. After digging a little further into the literature, it became apparent that DMF coordinates with the ruthenium metal center, effectively killing the catalytic activity.
Now you may be wondering where the residual DMF is actually coming from. The resin is washed several times with DCM, then DCE before adding the catalyst solution. In between each liquid addition though, the needle is washed with system solvent - in this case DMF. While the needles are fully cleaned in this process, there is residual DMF in the needle when the next liquid is removed from its storage container and through capillary action a small amount has been observed to essentially leak into the reagent container. For amino acids and other reagents already dissolved in DMF, this isn't a problem and the amount of extra DMF isn't sufficient to change the solution concentration.
In thinking about the options to deal with this poisoning issue two thoughts came to mind:
- -change the system solvent to DCM (or the like) after the synthesis is complete in preparation for the metathesis reaction
- -change catalyst load such that poisoning no longer occurs
The first option sounds awful from a practical standpoint and would likely require more time and energy than it is actually worth. Generally, the system solvent lines would have to be drained, bottle exchanged, and re-primed to equilibrate with DCM and then back to DMF for the final Fmoc deprotection before global deprotection and cleavage from the resin.
So I decided to increase the catalyst load, anticipating that a threshold would be achieved, enabling full conversion of the linear peptide to the desired stapled product. In the first series of experiments, I increased the catalyst load to 1-, 2-, or 3-fold relative to the literature reported levels, Figure 2.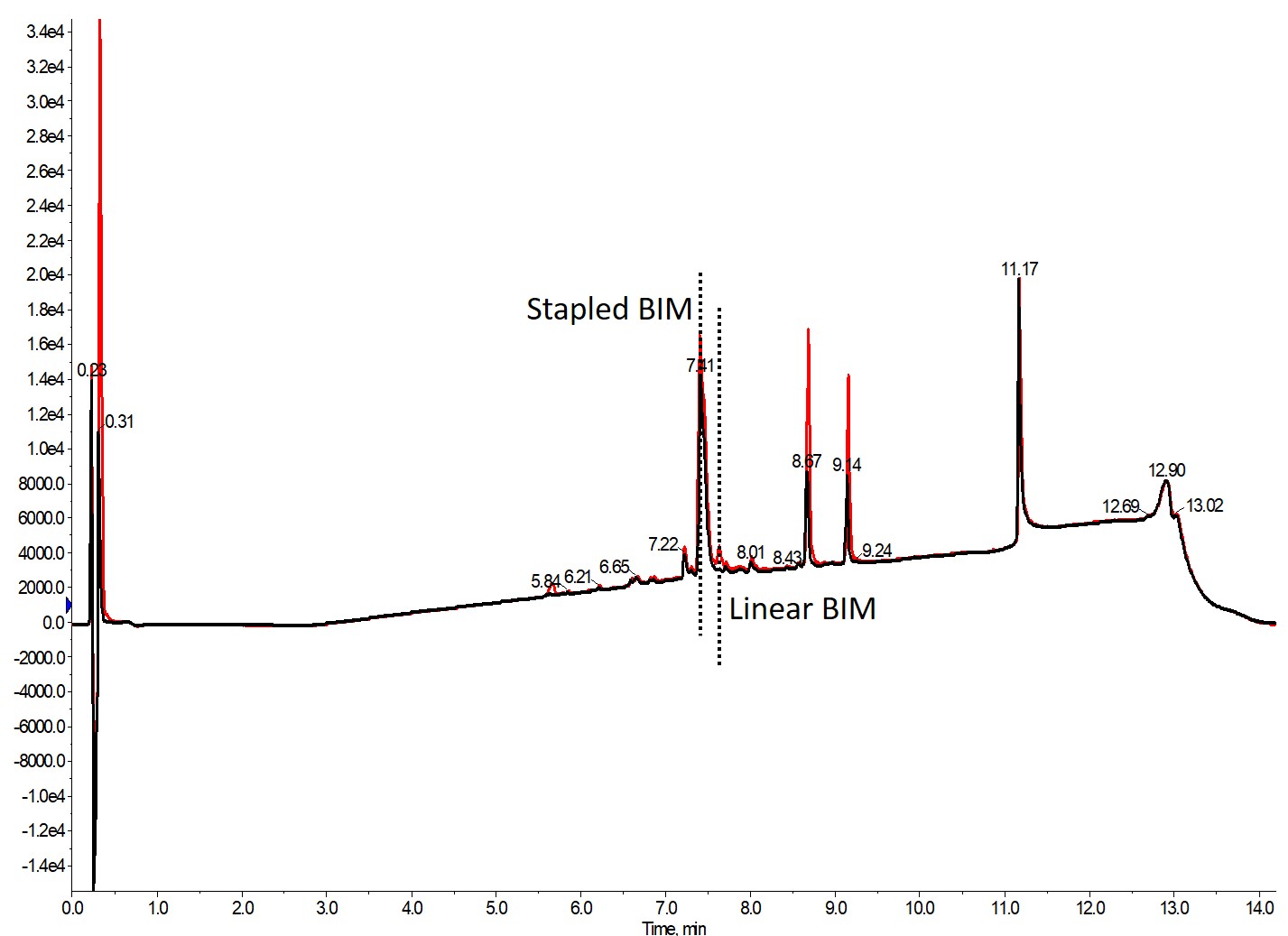
Figure 2. Reaction efficiency for RCM reaction comparing 20 mol percent (1 fold) added manually to the reactor vial or 40 mol percent (2 fold) Grubbs catalyst added automatically with the robot liquid handler. The reaction was allowed to proceed for 60 minutes at 40°C and only achieved 95% conversion with the 2-fold increase in catalyst load.
With 3-fold excess, or 60 mol percent catalyst load, full conversion to the stapled peptide product was finally achieved (data not shown). While this was ultimately the goal of this experiment, I started thinking about this whole process differently.
The Grubbs catalyst solution is prepared fresh before use rather than at the beginning of the synthesis to reduce exposure to the environment and avoid changes in concentration due to evaporation of the DCE solvent. Given this quick solution preparation, I decided that manually adding the catalyst solution was not so imposing when compared to the significant increase in reagent cost.
Although I would like to be able to tell you that fully automating everything is not only possible and should be achieved, there are certainly instances where a minimal manual intervention yields a more desirable result as in this case.
To learn more about how Biotage tools can help improve your laboratory efficiency, follow the link below.

 Organic Workflow
Organic Workflow Peptide Workflow
Peptide Workflow Scale-Up Flash Purification
Scale-Up Flash Purification  Sample Preparation
Sample Preparation Biomolecule Purification
Biomolecule Purification Oligo synthesis
Oligo synthesis Scavengers and Reagents
Scavengers and Reagents Service & Support
Service & Support Accessories & Spare parts
Accessories & Spare parts Investors
Investors Reports & News
Reports & News The Share
The Share Corporate Governance
Corporate Governance Calendar
Calendar Sustainability
Sustainability Our Offering
Our Offering Our History
Our History Our Locations
Our Locations Leadership
Leadership