I have experimented a lot with disulfide rich peptides lately, finding conditions that work well for not only the linear synthesis, but also for on-resin cysteine oxidations. Although simple scaffolds are useful for determining orthogonal protecting group removal and cysteine oxidation conditions, many of the peptides of interest today are much more complex – three or more disulfide bonds, and often head-to-tail cyclization.
In today’s post, I put to use three orthogonal protection strategies to optimize a fully automated synthesis of linaclotide.
Linaclotide, currently marketed for treatment of irritable bowl syndrome, is a 14 amino acid peptide that contains three disulfide bonds, Figure 1. This unique sequence presents complications in both the linear synthesis as well as with the disulfide bond pattern if a fully automated solution is desired.
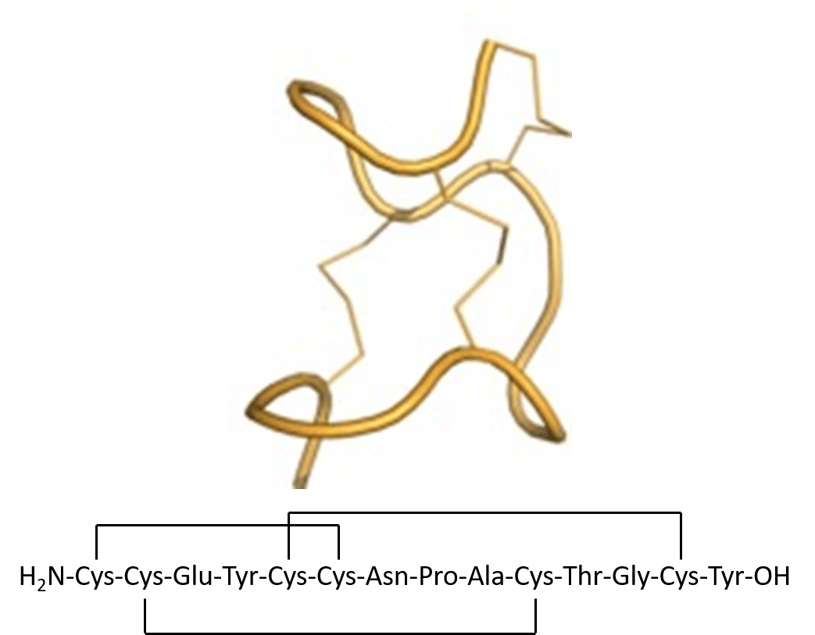
Figure 1. Solution structure and sequence of linaclotide.
To get started, I decided to use cysteine building blocks protected by Mmt, Acm and STmp protecting groups as I have worked with them individually in the past. But the most important question that quickly arose was which order to form the disulfide bonds. I’ve already talked about this some in the synthesis of apamin which contains two disulfide bonds. In my hands, the order of disulfide bond formation didn’t seem to matter for apamin, but that peptide is both longer in linear sequence and contains fewer disulfide bonds than linaclotide.
A quick note before we get into the actual syntheses – when using multiple orthogonal protecting groups it is important to consider compatibility of the removal conditions. For example, the Acm removal conditions are also sufficient to oxidize the Mmt protecting group which could potentially result in mixed disulfide bonds. Bearing this in mind, I’ll remove the STmp group first, then Mmt, and finally the Acm protecting group.For the synthesis, I used the Branches™ software feature on my Initiator+ Alstra™ to assign the order and associated chemistry for each of the regioselective disulfide bond reactions with confidence, Figure 2.
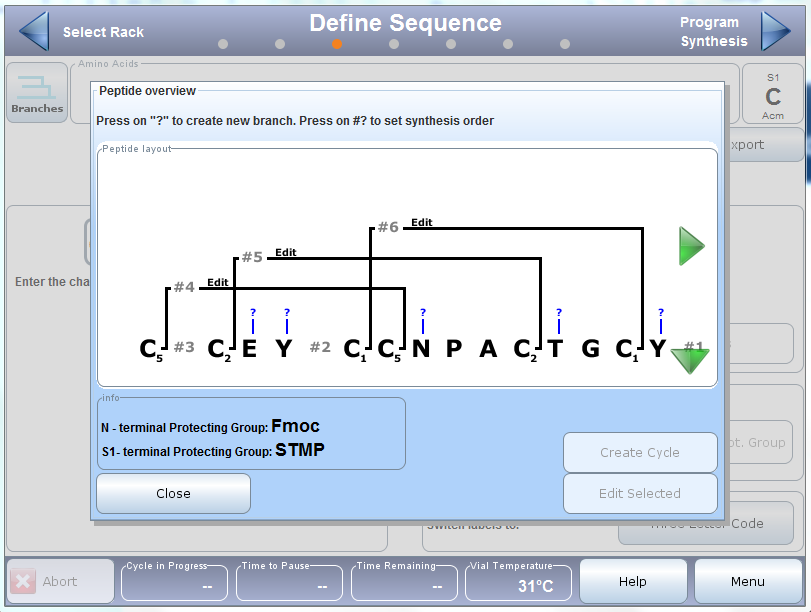
Figure 2. Branches representation of linaclotide sequence depicting not only the linear sequence and disulfide bond connectivity, but also the synthetic order.
I decided to oxidize the 5-13 disulfide bond first followed by 2-10 and finally the 1-6 disulfide bond, Figure 3.So, this didn’t work well at all. The broad underlying peak suggests the presence of multiple impurities, likely misfolded or incompletely oxidized forms of the desired peptide product. So back to the drawing board!
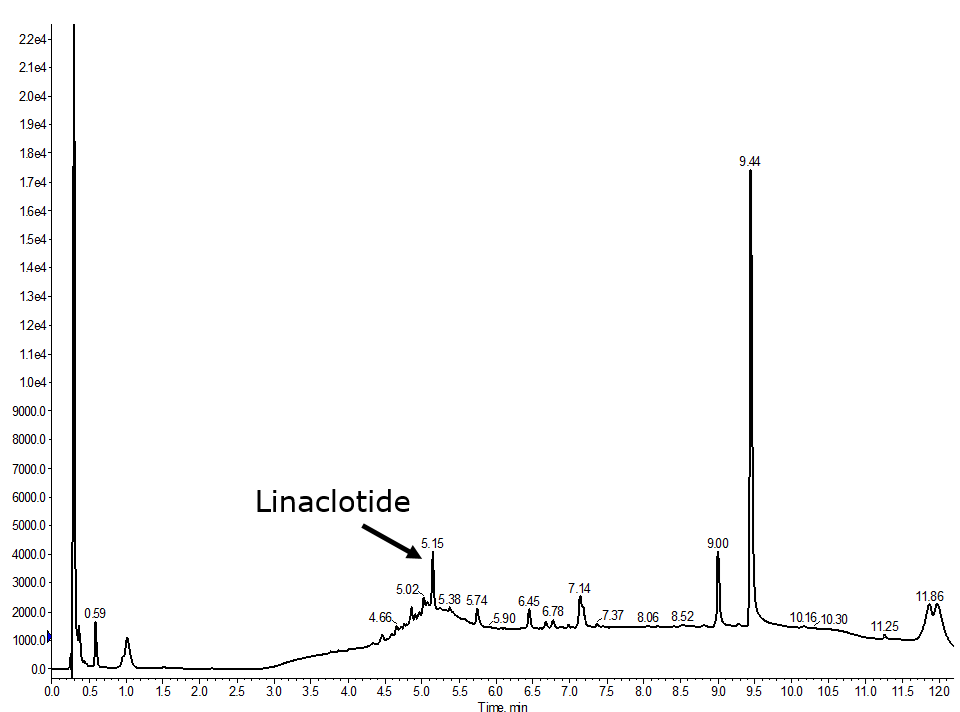
Figure 3. Analytical HPLC of a fully automated synthesis of linaclotide where the 3-15 disulfide bond was formed first, then 2-10 and finally the 1-6 was formed.
When the oxidation order is changed to the 1-6 first, followed by the 5-13 and finally the 2-10 disulfide bond (and no changes to the linear sequence synthesis protocol) things improve, Figure 4. Not only has the product peak intensity increased, the broad underlying peak has sharpened significantly, suggesting that the oxidation efficiency has improved.
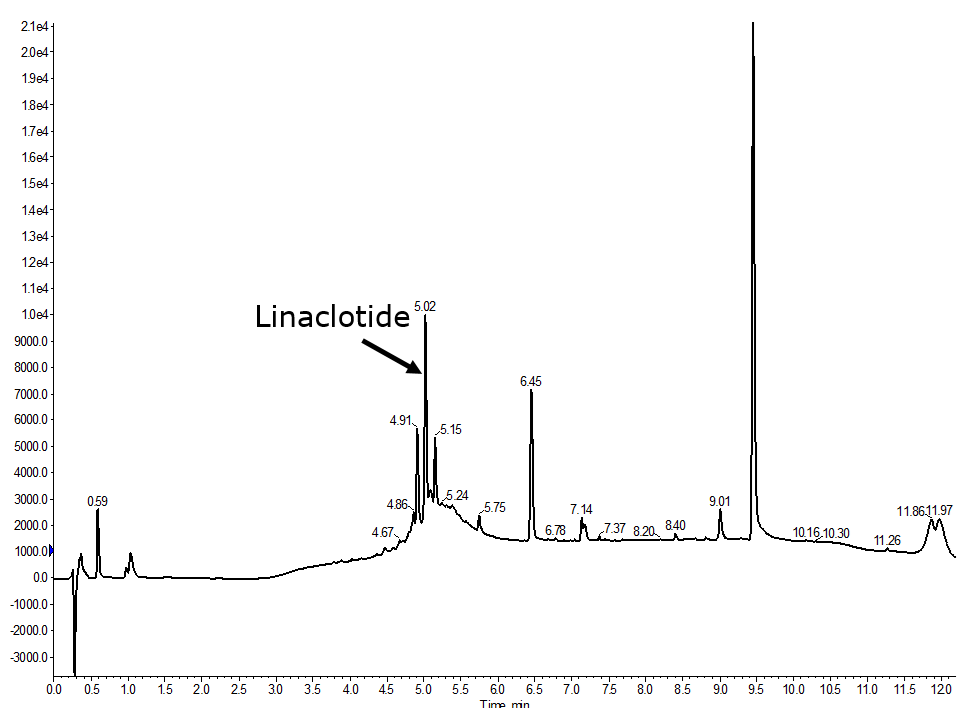
Figure 4. Analytical HPLC of a fully automated synthesis of linaclotide where the 1-6 disulfide bond was formed first, then 5-13, and finally the 2-10 was formed.
After taking a look at the literature (yes, I should have done this first) I found an article detailing an optimized synthesis of linaclotide and specifically details the order of disulfide bond formation. While they used a different series of protecting groups, the order of bond formation was most important for me. In a final synthesis, the 1-6 bond was formed first, followed by the 2-10 bond, and finally the 5-13 disulfide bond yielded the best results thus far, Figure 5.
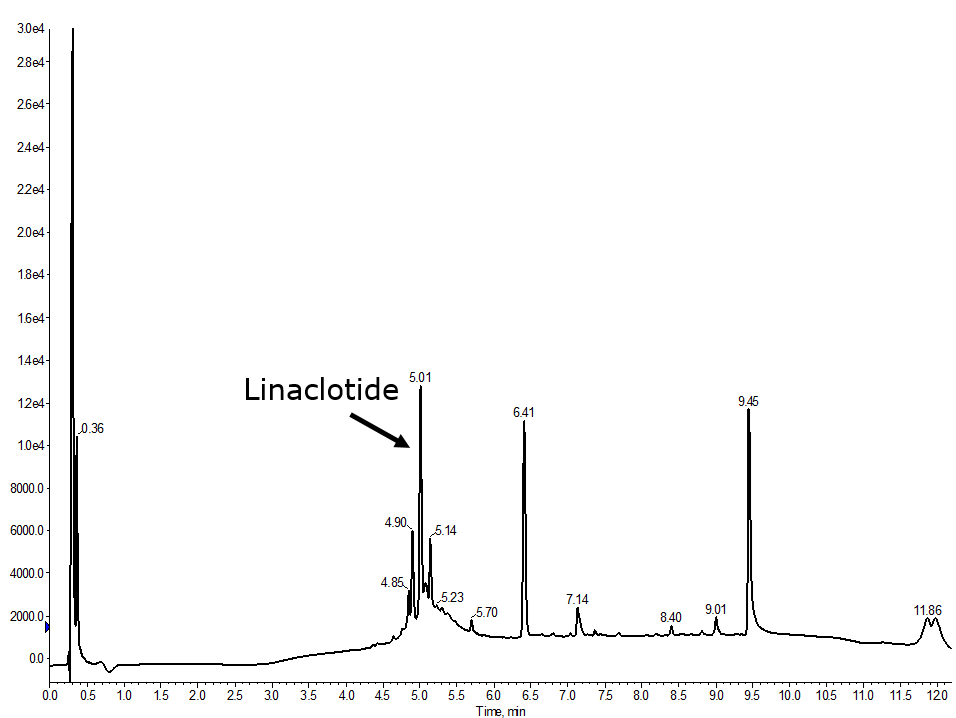
Figure 5. Analytical HPLC of a fully automated synthesis of linaclotide where the 1-6 disulfide bond was formed first, then 2-10, and finally the 5-13 was formed.
If we look at the solution structure of linaclotide, it makes some sense that this disulfide formation order is the most successful. The 5-13 bond, if formed first, could drive a conformation that would favor an interaction of the N-terminal region with the resin itself, hindering the oxidation efficiency of the other two disulfide bonds. Whereas, forming the 2-10 bond second facilitates the initial structural collapse, without the potential interaction with the resin polymer itself. Importantly, the 2-10 disulfide bonds also forms a structure that like positions the Cys-5 and Cys-13 for the final disulfide bond formation.
This process highlights the level of optimization that can be required to optimize synthesis automation for peptides that contain many disulfide bonds. Just think how much worse it could be for a peptide that adds in a head-to-tail cyclization!
If you want to learn more about synthesizing disulfide-rich peptides, watch my webinar by clicking the link below!

 Organic Workflow
Organic Workflow Peptide Workflow
Peptide Workflow Scale-Up Flash Purification
Scale-Up Flash Purification  Sample Preparation
Sample Preparation Biomolecule Purification
Biomolecule Purification Oligo synthesis
Oligo synthesis Scavengers and Reagents
Scavengers and Reagents Service & Support
Service & Support Accessories & Spare parts
Accessories & Spare parts Investors
Investors Reports & News
Reports & News The Share
The Share Corporate Governance
Corporate Governance Calendar
Calendar Sustainability
Sustainability Our Offering
Our Offering Our History
Our History Our Locations
Our Locations Leadership
Leadership