Hydrocarbon stapling as a strategy to stabilize secondary structures of peptides, while introduced by Miller, Blackwell and Grubbs in the mid 1990s, really grew to the forefront with seminal work by Schaffmeister and Verdine in early 2000s. Protocols have been developed that enable this post-synthesis modification while the peptide is still on resin, but often these metathesis reactions are performed manually, and at room temperature.
In today's post, I'll compare several different sets of reaction conditions using microwave heating with the goal of expediting the olefin metathesis reaction, without compromising reaction efficiency, and towards automating the entire synthesis.
Using peptides as tools to study protein-protein interactions and their related biological pathways just makes sense, especially with some structural data on hand. The two protein partners provide the natural template for which to build your tool compounds. Unfortunately though, many peptide fragments, when excised from their intact protein, lose all secondary structure and consequently, affinity for the target protein.
Many strategies for stabilizing a desired secondary structure, especially a helical structure, have been published. Hydrocarbon stapling, where two appropriately matched, non-natural amino acids each bearing a terminal alkene, can be reacted to form a non-native, covalent bridge across a linear peptide sequence.
The olefin metathesis chemistry pioneered by Grubbs and further applied to peptides by Verdine et al is extremely robust, but typically performed manually, after a linear synthesis is performed using an automated synthesizer. As a first step toward fully automated synthesis and metathesis, I decided to start with the BIM stapled peptide and explored some metathesis reaction conditions utilizing microwave heating.
I first synthesized the native linear BIM sequence, mostly to ensure that the basic synthesis strategy I chose would yield highly pure peptide and left the N-terminal Fmoc in place. Once confirmed, I used the same strategy, but this time incorporating the two olefin-bearing amino acids at positions 9 and 13 as published with reasonable success, Figure 1.
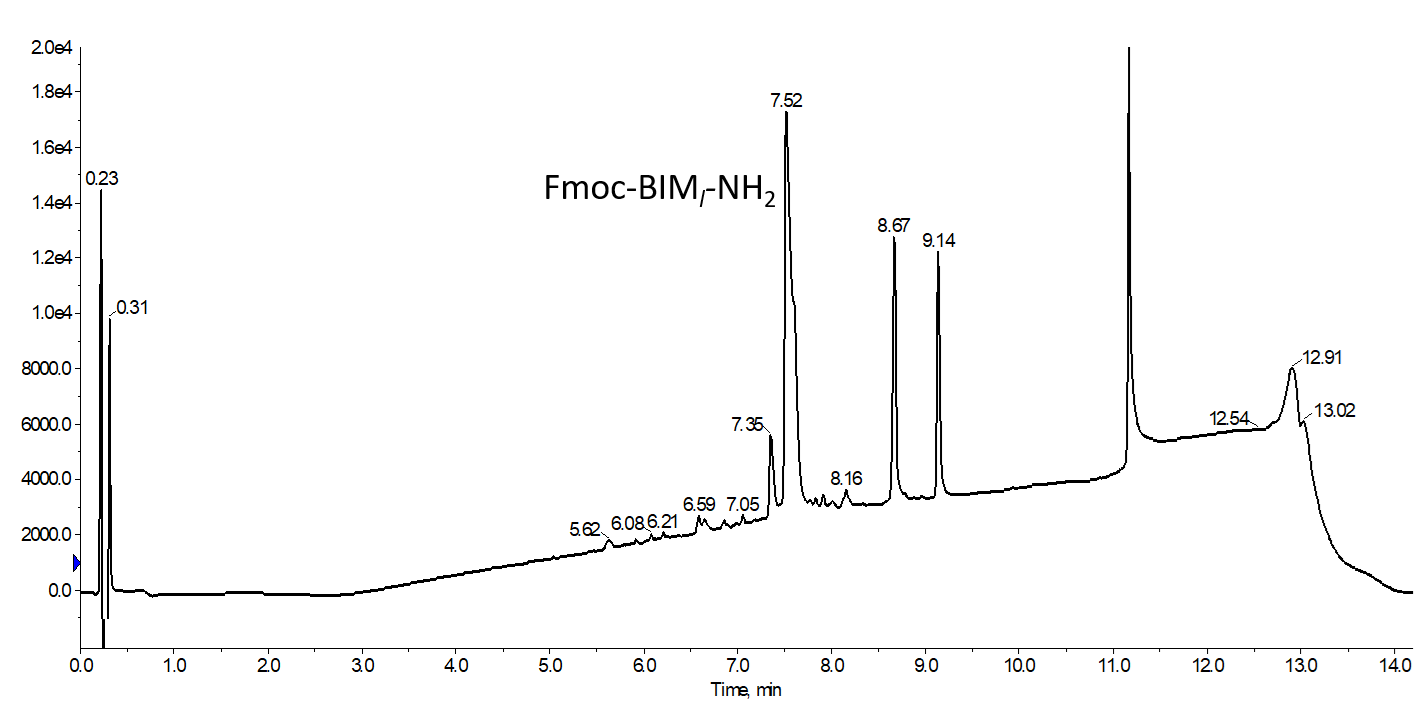
Figure 1. Analytical HPLC chromatogram for crude BIM peptide containing two unnatural, olefin-bearing amino acids prior to attempting any ring-closing metathesis reactions. The two large, late eluting peaks (8.67 and 9.14 min respectively) are attributed to residual protecting groups and show now peptidic masses in the mass spectrum.
I decided to start with the most standard protocol, performed manually as described, to confirm that this peptide would in fact behave as I expected (which it did), Figure 2.
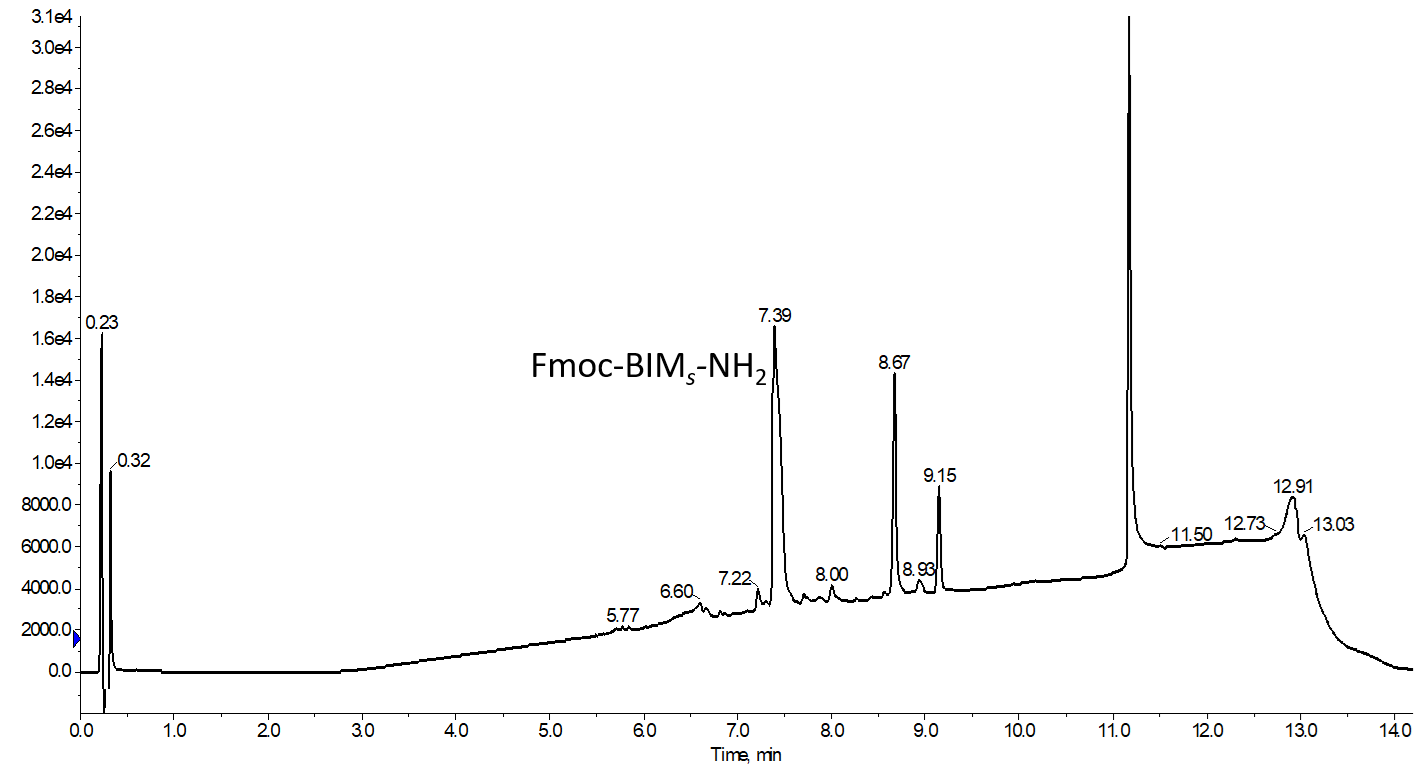
Figure 2. Analytical HPLC chromatogram of the BIM peptide stapled using the previously published, wholly manual procedure. Note the shift in retention time, commonly observed when comparing the linear and stapled peptides. There is no evidence for any remaining parent linear sequence either by UV or mass spec.
Before getting too far into the reaction conditions, it is important to note some concerns that may arise. The solvent used for the RCM reaction is 1,2-dichloroethane. It does have a higher boiling point than related dichloromethane, but both these solvents are nonpolar and therefore potentially difficult to heat with microwaves. The other principle concern is that the elevated temperature will cause the solvent to evaporate over the course of the reaction.
There is some precedent for microwave heating this reaction, so based on that, I used 40°C for the reaction, added the Grubbs catalyst solution manually, and then varied conditions as outlined in Table 1.
| Trial | Reaction Time (min) | Reaction Temp. | Reaction Iterations | Inert gas? | Percent Conversion |
| 1 | 120 | r.t | 2 | y | 100 |
| 2 | 30 | 40 | 2 | y | 100 |
| 3 | 30 | 40 | 2 | n | 100 |
| 4 | 30 | 40 | 1 | y | 95.0 |
| 5 | 30 | 40 | 1 | n | 78.5 |
| 6 | 60 | 40 | 1 | y | 99 |
| 7 | 60 | 40 | 1 | n | 96.8 |
Table 1. Ring-closing metathesis reaction conditions evaluated to convert the linear peptide sequence to the desired stapled peptide.
It isn't surprising to me that the conditions using two separate metathesis reaction iterations were successful. RCM, if performed properly is typically a highly efficient reaction. What was pleasantly surprising though is just how efficient a single reaction could be at elevated temperatures, even in the absence of an inert atmosphere.
It is important to note that the DCE solution absorbed microwave energy quite well and was easily heated throughout the course of the longest reactions evaluated here. Also, there was no significant evaporation observed with the open reactor vial that I used, even after heating for 60 minutes. This increased my confidence that this strategy can be effective for a wide variety of stapled peptides.
Inert gas is not required for the general operation of the Biotage® Initator+ Alstra™, but the instrument can be equipped with an inert gas manifold, protecting not only the amino acid and reagents solutions, but also purging the microwave cavity during reactions. It is interesting to me the difference in conversion efficiency when comparing a set of reaction conditions either with or without the inert gas (nitrogen in this case) purging the microwave cavity, Figure 3.
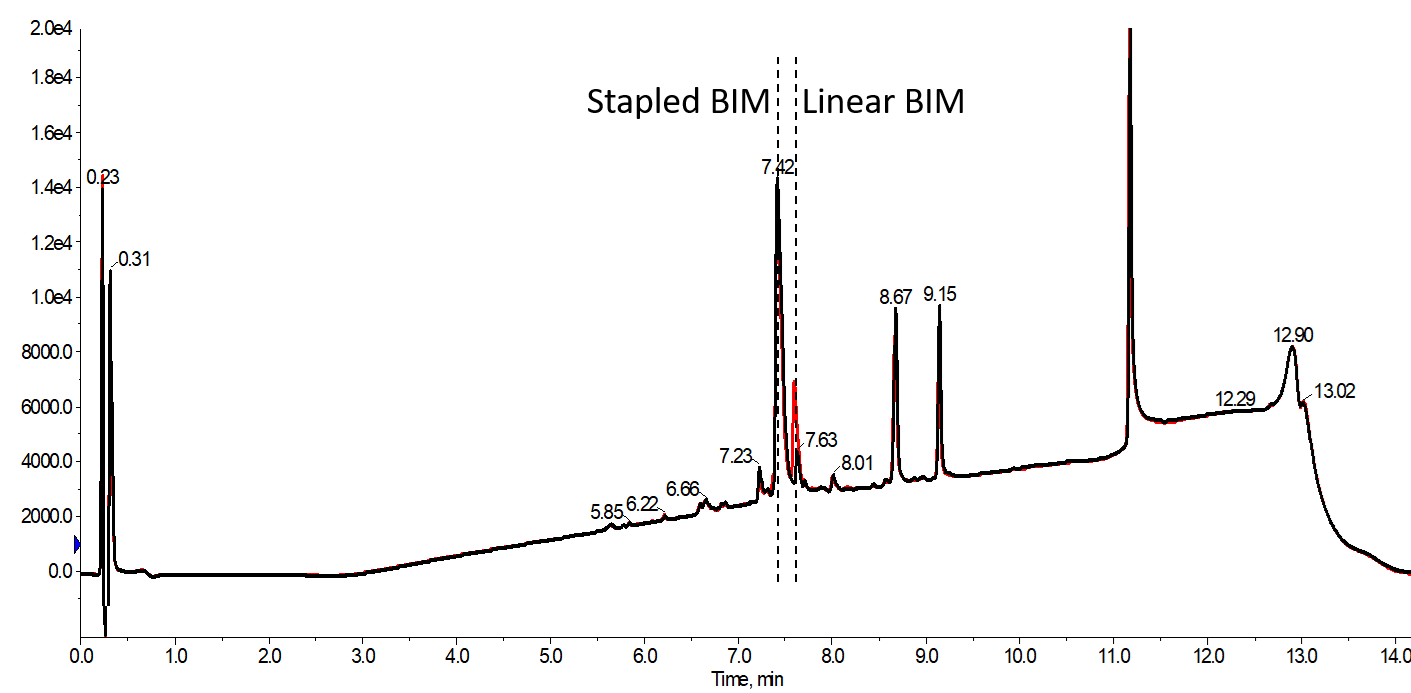
Figure 3. Analytical HPLC chromatograms for peptides stapled for 30 min at 40 °C using only one RCM reaction either with (black) or without (red) inert gas purging the microwave cavity during the reaction. The yield of desired, cyclized BIM peptide decreases to only 78% simply by removing the inert gas. This is difference is also observed for a single RCM reaction for 60 min, but to a lesser extent.
There are two possibilities distinct for this difference:
-The Grubbs catalyst is degrading due to exposure to the environment. It's well established that the first generation catalyst is somewhat sensitive to environmental oxygen.
-Nitrogen bubbling used to mix the resin slurry during the "standard" reaction protocol is driving off the ethylene byproduct, further driving the metathesis reaction to completion.
My guess at this point is that this is likely the result of a combination of both these phenomena at this point. I'll be testing out the first one in some future work with the 2nd generation catalyst.
To learn more about how Biotage tools can improve the efficiency of peptide synthesis, follow the link below.

 Organic Workflow
Organic Workflow Peptide Workflow
Peptide Workflow Scale-Up Flash Purification
Scale-Up Flash Purification  Sample Preparation
Sample Preparation Biomolecule Purification
Biomolecule Purification Oligo synthesis
Oligo synthesis Scavengers and Reagents
Scavengers and Reagents Service & Support
Service & Support Accessories & Spare parts
Accessories & Spare parts Investors
Investors Reports & News
Reports & News The Share
The Share Corporate Governance
Corporate Governance Calendar
Calendar Sustainability
Sustainability Our Offering
Our Offering Our History
Our History Our Locations
Our Locations Leadership
Leadership