Have you ever wondered if there was a faster (and possibly cheaper) way to purify your peptides?
My colleagues and I in the peptide community rely almost exclusively on reversed-phase HPLC for delivery of highly pure peptide products. However, this process is often very time consuming and requires expensive columns and solvents to be successful. Alternatively, peptide purification via reversed-phase flash column chromatography can be used to complete a purification in a fraction of the time and with a fraction of the costs.
Here I will show how I do gradient optimization for peptide purification via reversed-phase flash column chromatography and will highlight the similarities with standard HPLC methodologies.
While I have had the privilege of purifying peptides with a fully automated mass-directed HPLC system, I have also purified peptides the old fashioned, painful way manually injecting sample and manually collecting fractions. Now that I’ve started my career with Biotage and have been exposed to flash column chromatography, I decided to test out whether the rules for gradient optimization in standard HPLC can also be applied to a flash column chromatography.
So, I synthesized a 28-amino acid peptide on a 200 µmol scale automatically with a Biotage® Initiator+ Alstra™ peptide synthesizer and recovered approximately 400 mg of crude product. My peptide of interest was in fact present as determined analytical HPLC-MS, which showed that the peptide elutes in 25%-30% acetonitrile. At this point, I used to set up an HPLC with an autosampler and begin the process of multiple (often many) injections of the crude peptide mixture. This time though, I switched to a reversed-phase flash chromatography system to purify my peptide.
I first dissolved my peptide in DMSO to enable maximum loading onto the column in a minimal volume. If you are concerned about oxidation of Cys or Met residues during the purification, try DMF as an alternative high-capacity injection solvent.
I decided to load 50 mg of crude sample onto a 12 g Biotage® Sfär Ultra C18 cartridge while I identified an optimal mobile phase gradient. The 50 mg load corresponds to 0.4% of the amount of C18 in the cartridge, a 4-fold increase in column loading over a typical RP-HPLC column.
For my first injection, the default reversed-phase gradient was slightly modified by increasing the flow rate from 12 mL/min to 50 mL/min, Figure 1. The higher flow rate minimizes peak decomposition due to the pump’s action while staying within the confines of the cartridge’s pressure limitations. The gradient was set from 10% to 100% acetonitrile over 10 column volumes (6.36% per min) for two reasons. The first was to ensure that the peptide elution profile is similar to that observed when using my analytical HPLC column. (Not all C18 columns are created equal and it is well established that particle size, pore size, and silanol chemistry can dramatically impact molecular retention.) The second reason is that I wanted to establish a framework for the subsequent gradients to be evaluated. It is difficult to compare separation efficiency when the length of the gradient changes, therefore I kept the gradient length constant.
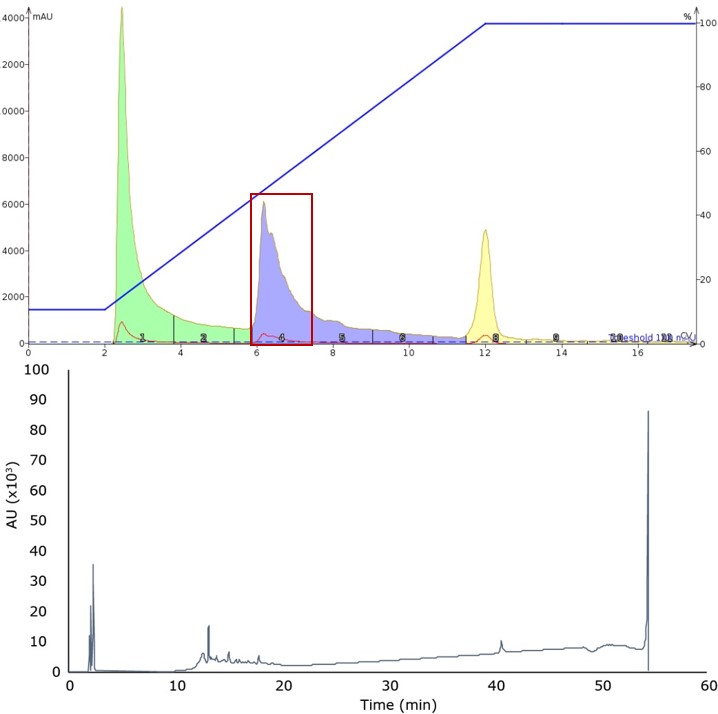
Figure 1. Peptide elution profile observed during the initial gradient, covering 10% to 100% acetonitrile (top) and analytical HPLC chromatogram of the predominant fraction (bottom). The desired product was identified in the main peak (approximately 12 min retention time) by mass spectrometry.
While the peak shape is relatively sharp, the trailing shoulder suggests some other compounds are present in the sample. To further resolve and isolate the contaminating peptide fragments, I decreased the slope of the gradient, running from 20% to 60% acetonitrile over 10 CV (2.82% per min), Figure 2.
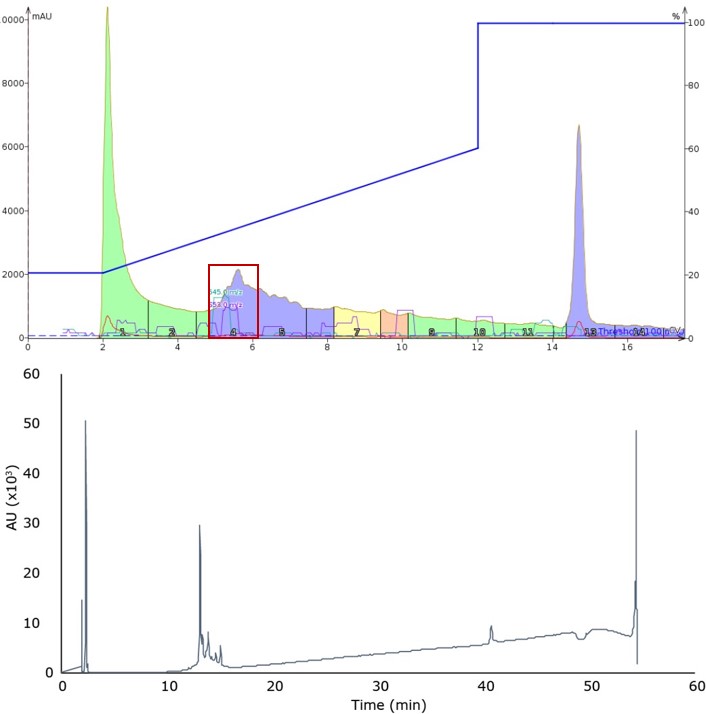
Figure 2. Peptide elution profile observed during the second gradient covering 20% to 60% acetonitrile (top) and analytical HPLC of the main peak, fraction 4 (bottom).
Though the original peak shape has clearly worsened, it resulted in an improvement in separation and overall fraction purity, indicated by comparing the analytical HPLC-MS for the main peak fraction isolated during injection 1 and injection 2, Figure 2. This suggests that using a shallow gradient with high flow rate can improve peptide sample purity with flash column chromatography just as in standard RP-HPLC.
For the third injection, I decreased the gradient slope even further, reducing the change in mobile phase concentrations to a rate similar to that utilized in HPLC. Although this gradient, from 20% to 30% acetonitrile over 10 CV (0.706% per min) generated the ugliest chromatography of the group thus far, it also yielded the cleanest fractions containing the desired peptide product, Figure 3.
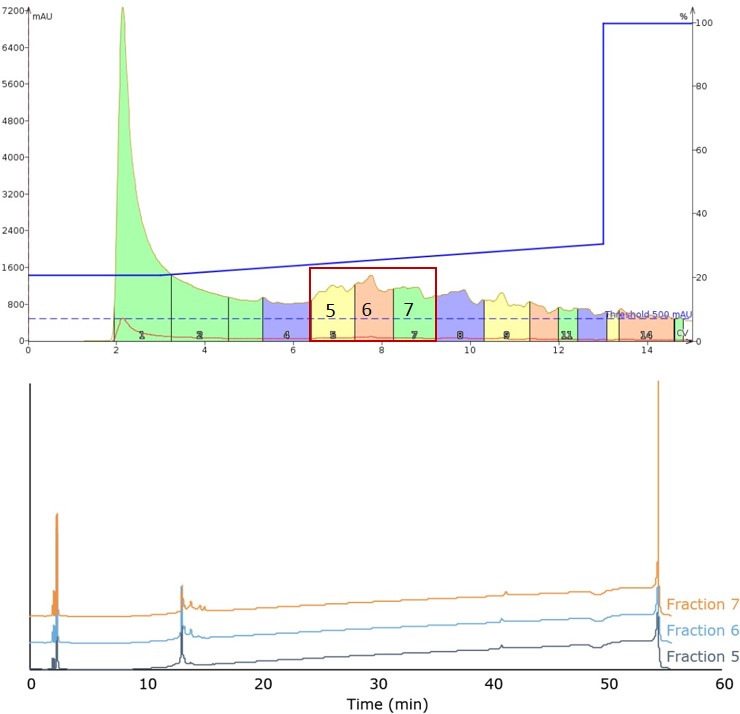
Figure 3. Peptide elution of the third mobile phase gradient, covering 20% to 30% acetonitrile (top). Peak broadening required analytical HPLC analysis of fraction 5, fraction 6, and fraction 7 (bottom). The main peak contains the desired product in all fractions.
With this optimized gradient in hand, I sought to purify the remaining 200 mg of peptide as rapidly as possible. Following the rules that are typically applied for linear scaling a flash chromatography method, I chose a 60 g SNAP Ultra C18 cartridge, increased the flow rate to maintain the linear velocity, and injected all 200 mg onto the cartridge in 800 µL of DMSO, Figure 4. It is important to note that the crude sample, even at significantly larger injection volume, retained its chromatographic behavior. This demonstrates that the gradient optimized at smaller scale was easily transferred to a larger scale purification.
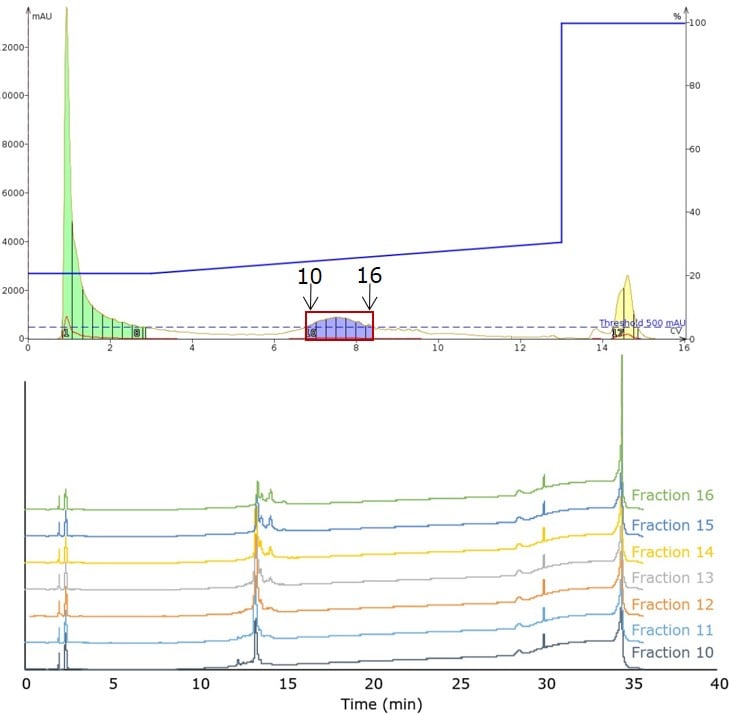
Figure 4. Peptide elution profile observed during the large scale injection (200 mg) and purification using the optimized mobile phase gradient (top). Analytical HPLC evaluation of each fraction contained within the main peak (bottom).
Evaluating the analytical HPLC trace for each collected fraction shows the purity that can be rapidly achieved in peptide purification by flash column chromatography. The first several fractions are potentially suitable for immediate use, while the later fractions will require an HPLC pass to reach appropriate purity levels, as expected from the previous injections. I can also improve the first-pass recovery of pure peptide by reducing the volume of each fraction collected.
It is important to note that each scouting run was completed in approximately 25 minutes, including column equilibration and washing and the majority of the crude sample mixture was purified in less than 1 hour.
There are several key conclusions to be drawn from gradient evaluation:
1) standard HPLC techniques for gradient optimization are readily transferable to flash column chromatography
2) the rules for scaling sample loading can be applied to reversed-phased methods for peptide purification
3) large quantities of crude peptide sample can be rapidly purified, significantly reducing the time required to fully purify a crude peptide sample.
Have you thought about using flash chromatography to purify your peptide samples? Follow the link below to learn different strategies to achieve highly pure peptides.

 Organic Workflow
Organic Workflow Peptide Workflow
Peptide Workflow Scale-Up Flash Purification
Scale-Up Flash Purification  Sample Preparation
Sample Preparation Biomolecule Purification
Biomolecule Purification Oligo synthesis
Oligo synthesis Scavengers and Reagents
Scavengers and Reagents Service & Support
Service & Support Accessories & Spare parts
Accessories & Spare parts Investors
Investors Reports & News
Reports & News The Share
The Share Corporate Governance
Corporate Governance Calendar
Calendar Sustainability
Sustainability Our Offering
Our Offering Our History
Our History Our Locations
Our Locations Leadership
Leadership